-
Notifications
You must be signed in to change notification settings - Fork 42
/
27-dimension-reduction.qmd
618 lines (427 loc) · 21.8 KB
/
27-dimension-reduction.qmd
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
148
149
150
151
152
153
154
155
156
157
158
159
160
161
162
163
164
165
166
167
168
169
170
171
172
173
174
175
176
177
178
179
180
181
182
183
184
185
186
187
188
189
190
191
192
193
194
195
196
197
198
199
200
201
202
203
204
205
206
207
208
209
210
211
212
213
214
215
216
217
218
219
220
221
222
223
224
225
226
227
228
229
230
231
232
233
234
235
236
237
238
239
240
241
242
243
244
245
246
247
248
249
250
251
252
253
254
255
256
257
258
259
260
261
262
263
264
265
266
267
268
269
270
271
272
273
274
275
276
277
278
279
280
281
282
283
284
285
286
287
288
289
290
291
292
293
294
295
296
297
298
299
300
301
302
303
304
305
306
307
308
309
310
311
312
313
314
315
316
317
318
319
320
321
322
323
324
325
326
327
328
329
330
331
332
333
334
335
336
337
338
339
340
341
342
343
344
345
346
347
348
349
350
351
352
353
354
355
356
357
358
359
360
361
362
363
364
365
366
367
368
369
370
371
372
373
374
375
376
377
378
379
380
381
382
383
384
385
386
387
388
389
390
391
392
393
394
395
396
397
398
399
400
401
402
403
404
405
406
407
408
409
410
411
412
413
414
415
416
417
418
419
420
421
422
423
424
425
426
427
428
429
430
431
432
433
434
435
436
437
438
439
440
441
442
443
444
445
446
447
448
449
450
451
452
453
454
455
456
457
458
459
460
461
462
463
464
465
466
467
468
469
470
471
472
473
474
475
476
477
478
479
480
481
482
483
484
485
486
487
488
489
490
491
492
493
494
495
496
497
498
499
500
501
502
503
504
505
506
507
508
509
510
511
512
513
514
515
516
517
518
519
520
521
522
523
524
525
526
527
528
529
530
531
532
533
534
535
536
537
538
539
540
541
542
543
544
545
546
547
548
549
550
551
552
553
554
555
556
557
558
559
560
561
562
563
564
565
566
567
568
569
570
571
572
573
574
575
576
577
578
579
580
581
582
583
584
585
586
587
588
589
590
591
592
593
594
595
596
597
598
599
600
601
602
603
604
605
606
607
608
609
610
611
612
613
614
615
616
617
618
# Dimension reduction
```{r, echo=FALSE}
rafalib::mypar()
```
* General idea is to reduce the dimension of the dataset while preserving important characteristics, such as the distance between features or observations.
* With fewer dimensions, visualization then becomes more feasible.
* The technique behind it all, the singular value decomposition, is also useful in other contexts. Principal component analysis (PCA) is the approach we will be showing.
* We will motivate the ideas behind with a simple example.
## Motivation: preserving distance
* We simulated heights of pairs of twins, some adults some children.
* We use the `mvrnorm` function from the **MASS** package to simulate bivariate normal data.
```{r, message=FALSE}
set.seed(1983)
library(MASS)
n <- 100
Sigma <- matrix(c(9, 9 * 0.9, 9 * 0.9, 9), 2, 2)
x <- rbind(mvrnorm(n / 2, c(69, 69), Sigma),
mvrnorm(n / 2, c(60, 60), Sigma))
```
* A scatterplot quickly reveals that the correlation is high and that there are two groups of twins, the adults (upper right points) and the children (lower left points):
```{r distance-illustration, fig.width=3, fig.height=3, echo=FALSE, out.width="40%"}
rafalib::mypar()
lim <- range(x)
plot(x, xlim = lim, ylim = lim)
lines(x[c(1, 2),], col = "blue", lwd = 2)
lines(x[c(2, 51),], col = "red", lwd = 2)
points(x[c(1, 2, 51),], pch = 16)
```
* We want to explore the data through a histogram of a one-dimensional variable.
* We want to reduce the dimensions from two to one, but still be able to understand important characteristics of the data, for example that the observations cluster into two groups: adults and children.
* To show the ideas presented here are generally useful, we will standardize the data so that observations are in standard units rather than inches:
```{r}
x <- scale(x)
```
In the figure above we show the distance between observation 1 and 2 (blue), and observation 1 and 51 (red). Note that the blue line is shorter, which implies 1 and 2 are closer.
We can compute these distances using `dist`:
```{r}
d <- dist(x)
as.matrix(d)[1, 2]
as.matrix(d)[2, 51]
```
This distance is based on two dimensions and we need a distance approximation based on just one.
Let's start with the naive approach of simply removing one of the two dimensions. Let's compare the actual distances to the distance computed with just the first dimension:
```{r}
z <- x[,1]
```
To make the distances comparable, we divide the sum of squares by the number of dimensions. So for the two dimensional case we have
$$
\sqrt{ \frac{1}{2} \sum_{j=1}^2 (x_{1,j}-x_{2,j})^2 },
$$
so we divide the distance by $\sqrt{2}$:
```{r, fig.width=3, fig.height=3, out.width="40%"}
plot(dist(x) / sqrt(2), dist(z))
abline(0, 1, col = "red")
```
* Based on the plot, this one number summary does ok at preserving distances, but, can we pick a one-dimensional summary that makes this one-number approximation even better?
## Rotations
Any two-dimensional point $(X_1, X_2)$ can be written as the base and height of a triangle with a hypotenuse going from $(0,0)$ to $(X_1, X_2)$:
$$
x_1 = r \cos\phi, \,\, x_2 = r \sin\phi
$$
with $r$ the length of the hypothenus and $\phi$ the angel between the hypotenuse and the x-axis.
We can *rotate* the point $(x_1, x_2)$ around a circle with center $(0,0)$ and radius $r$ by an angle $\theta$ by changing the angle in the previous equation to $\phi + \theta$:
$$
z_1 = r \cos(\phi+ \theta), \,\,
z_2 = r \sin(\phi + \theta)
$$
```{r, echo=FALSE, fig.asp=0.7}
draw.circle <- function(angle, start = 0, center = c(0,0), r = 0.25){
th <- seq(start, start + angle, length.out = 100)
x <- center[1] + r*cos(th)
y <- center[2] + r*sin(th)
lines(x, y)
}
rafalib::mypar()
rafalib::nullplot(-0.25, 1.75,-0.25, 1.05, axes = FALSE)
abline(h=0,v=0, col= "grey")
draw.circle(2*pi, r=1)
phi <- pi/12
arrows(0, 0, 0.975*cos(phi), 0.975*sin(phi), length = 0.1)
points(cos(phi), sin(phi), pch = 16)
text(0.3*cos(phi/2), 0.3*sin(phi/2), expression(phi), font = 2)
text(cos(phi), sin(phi), expression('(' * x[1] * ',' * x[2] * ') = (' * phantom(.) * 'r' * phantom(.) * 'cos('*phi*'), r' * phantom(.) * 'sin('*phi*')' * phantom(.) * ')' ),
pos=4)
draw.circle(phi)
theta <- pi/4
points(cos(phi+ theta), sin(phi+theta), pch = 16)
arrows(0, 0, 0.975*cos(phi+theta), 0.975*sin(phi+theta), length = 0.1)
text(0.35*cos(phi+theta/2), 0.35*sin(phi+theta/2), expression(theta), font = 2)
text(cos(phi + theta), sin(phi + theta),
expression('(' * z[1] * ',' * z[2] * ') = (' * phantom(.) * 'r' * phantom(.) * 'cos('*phi*'+'*theta*'), r' * phantom(.) * 'sin('*phi*'+'*theta*')' * phantom(.) * ')' ), pos = 4)
draw.circle(theta, start = phi, r = 0.3)
```
We can use trigonometric identities to rewrite $(z_1, z_2)$ in the following way:
$$
\begin{align}
z_1 = r \cos(\phi + \theta) = r \cos \phi \cos\theta - r \sin\phi \sin\theta = x_1 \cos(\theta) - x_2 \sin(\theta)\\
z_2 = r \sin(\phi + \theta) = r \cos\phi \sin\theta + r \sin\phi \cos\theta = x_1 \sin(\theta) + x_2 \cos(\theta)
\end{align}
$$
* Now we can rotate each point in the dataset by simply applying the formula above to each pair $(x_{i,1}, x_{i,2})$.
* Here is what the twin standardized heights look like after rotating each point by $-45$ degrees:
```{r, fig.width = 6, fig.height = 3, echo=FALSE}
z <- cbind((x[,1] + x[,2]) / sqrt(2), (x[,2] - x[,1]) / sqrt(2))
lim <- range(z)
rafalib::mypar(1,2)
plot(x, xlim=lim, ylim = lim)
lines(x[c(1,2),], col = "blue", lwd = 2)
lines(x[c(2,51),], col = "red", lwd = 2)
points(x[c(1,2,51),], pch = 16)
plot(z, xlim=lim, ylim = lim)
lines(z[c(1,2),], col = "blue", lwd = 2)
lines(z[c(2,51),], col = "red", lwd = 2)
points(z[c(1,2,51),], pch = 16)
```
* Note that while the variability of $x_1$ and $x_2$ are similar, the variability of $z_1$ is much larger than the variability of $z_2$.
* Also note that the distances between points appear to be preserved.
## Linear transformations
* Note that each row of $\mathbf{X}$ was transformed using a linear transformation.
* For any row $i$, the first entry was:
$$z_{i,1} = a_{11} x_{i,1} + a_{21} x_{i,2}$$
with $a_{11} = \cos\theta$ and $a_{21} = -\sin\theta$.
* The second entry was also a linear transformation:
$$z_{i,2} = a_{12} x_{i,1} + a_{22} x_{i,2}$$
with $a_{12} = \sin\theta$ and $a_{22} = \cos\theta$.
* We can also use linear transformations to get $X$ back from $Z$.
* Solving the system of two linear equations gives us:
$$x_{i,1} = b_{1,1} z_{i,1} + b_{2,1} z_{i,2}$$
with $b_{1,2} = \cos\theta$ and $b_{2,1} = \sin\theta$
and
$$x_{i,2} = b_{2,1} z_{i,1} + b_{2,2} z_{i,2}$$
with $b_{2,1} = -\sin\theta$ and $b_{1,2} = \cos\theta$.
* Using linear algebra, we can write the operation we just performed like this:
$$
\begin{pmatrix}
z_1&z_2
\end{pmatrix}
=
\begin{pmatrix}
x_1&x_2
\end{pmatrix}
\begin{pmatrix}
a_{11}&a_{12}\\
a_{21}&a_{22}
\end{pmatrix}
$$
* An advantage of using linear algebra is that we can write the transformation for the entire dataset by representing it as a $N \times 2$ matrix $X$, with each row holding the two values for a pair of twins, and the rotation as a *linear transformation* of $X$:
$$
\mathbf{Z} = \mathbf{X} \mathbf{A}
\mbox{ with }
\mathbf{A} = \,
\begin{pmatrix}
a_{11}&a_{12}\\
a_{21}&a_{22}
\end{pmatrix}.
$$
* This transformation results in an $N \times 2$ matrix, denoted here with $Z$, with the rotated points in each row.
* Another advantage of linear algebra is that we can rotate back by simply multiplying $Z$ by the inverse matrix:
$$
\mathbf{Z} \mathbf{A}^{-1} = \mathbf{X} \mathbf{A} \mathbf{A}^{-1} = \mathbf{X}
$$.
* This implies that all the information in $\mathbf{X}$ is included in the rotation $\mathbf{Z}$, and it can be retrieved via a linear transformation.
* These derivations imply that we can use the following code to rotate the data by any angle $\theta$:
```{r}
rotate <- function(x, theta){
theta <- theta/360 * 2 * pi # convert to radians
A <- matrix(c(cos(theta), -sin(theta), sin(theta), cos(theta)), 2, 2)
x %*% A
}
```
* We can use this to confirm that for any rotation the distances are preserved. Here are two examples:
```{r}
max(dist(rotate(x, -45)) - dist(x))
max(dist(rotate(x, 30)) - dist(x))
```
## Orthogonal transformations
* Recall that the distance between two points, say rows $h$ and $i$ of the transformation $\mathbf{Z}$, can be written like this:
$$
||\mathbf{z}^\top_h - \mathbf{z}^\top_i|| = (\mathbf{z}_h - \mathbf{z}_i)(\mathbf{z}^\top_h - \mathbf{z}^\top_i)
$$
with $\mathbf{z}_h$ and $\mathbf{z}_i$ the $h$-th and $i$-th $1 \times p$ rows, with $p=2$ in our specific example.
* Using linear algebra, we can rewrite this quantity as
$$
||\mathbf{z}^\top_h - \mathbf{z}^\top_i||^2 = ||\mathbf{A}^\top\mathbf{x}^\top_h - \mathbf{A}^\top\mathbf{x}^\top_i||^2 = (\mathbf{x}_h - \mathbf{x}_i) \mathbf{A} \mathbf{A}^\top (\mathbf{x}^\top_h - \mathbf{x}^\top_i)
$$
* Note that if $\mathbf{A} \mathbf{A}^\top = \mathbf{I}$ then the distance between the $h$th and $i$th rows is the same for the original and transformed data.
* We to transformation with the property $\mathbf{A} \mathbf{A}^\top = \mathbf{I}$ as _orthogonal transformations_ and they are guaranteed to preserves the distance between any two points.
* Notice that $\mathbf{A}$ being orthogonal also guarantees that the total sum of squares of any row in $\mathbf{X}$ is preserved after the transformation:
$$
||\mathbf{z}^\top_i||^2 = ||\mathbf{x}^\top_i\mathbf{A}||^2 = \mathbf{x}_i \mathbf{A}\mathbf{A}^\top \mathbf{x}^\top_i = \mathbf{x}_i \mathbf{x}^\top_i = ||\mathbf{x}^\top_i||^2
$$
* Our transformation has this property:
```{r}
theta <- 30 # works for any theta
A <- matrix(c(cos(theta), -sin(theta), sin(theta), cos(theta)), 2, 2)
A %*% t(A)
```
* We can confirm this by computing the total variation for `x` and `z`:
```{r}
z <- rotate(x, -45)
sum(colSums(x^2))
sum(colSums(z^2))
```
* This can be interpreted as consequence of the fact that orthogonal transformation guarantee that all the information is preserved.
## Principal Component Analysis (PCA) {#sec-pca}
* Now how does this all relate to our goal of finding a one-dimensional summary that approximates distance between points?
* Note that in our original data the variability of the two dimensions is the same:
```{r}
colSums(x^2)
```
* But after a rotations this is no longer true:
```{r}
colSums(z^2)
```
* With `z`, the proportion of the variability included in the first dimension is much higher.
* We can search for the rotation that maximizes the proportion of the variability included in the first dimension:
```{r max-rotation, fig.width=3, fig.height=3, echo=FALSE, out.width="50%"}
library(tidyverse)
rafalib::mypar()
angles <- seq(0, -90)
v <- sapply(angles, function(angle) colSums(rotate(x, angle)^2))
variance_explained <- v[1,] / (v[1,] + v[2,])
plot(angles, variance_explained)
```
```{r, eval=FALSE}
angles <- seq(0, -90)
v <- sapply(angles, function(angle) colSums(rotate(x, angle)^2))
variance_explained <- v[1,] / (v[1,] + v[2,])
plot(angles, variance_explained)
```
* We see that the variability included in the first dimension is maximized at about -45 degrees.
* Becuase almost all the variation is explained by this first dimension, with this particular rotation the distance between points in $X$ can be very well approximated by just the first dimension of $Z$, much better than with the first dimension of $X$:
```{r distance-approx-2, echo=FALSE, fig.width=3, fig.height=3, out.width="40%"}
rafalib::mypar()
plot(dist(x), dist(z[,1]))
abline(0,1, col = "red")
```
* We also notice that the two groups, adults and children, can be clearly observed with the one number summary:
```{r}
hist(z[,1], nclass = 15)
```
Below, we learn that the rotation that maximizes the standard deviation of the first dimention `z[,1]` gives us the first principal component of the matrix `x`.
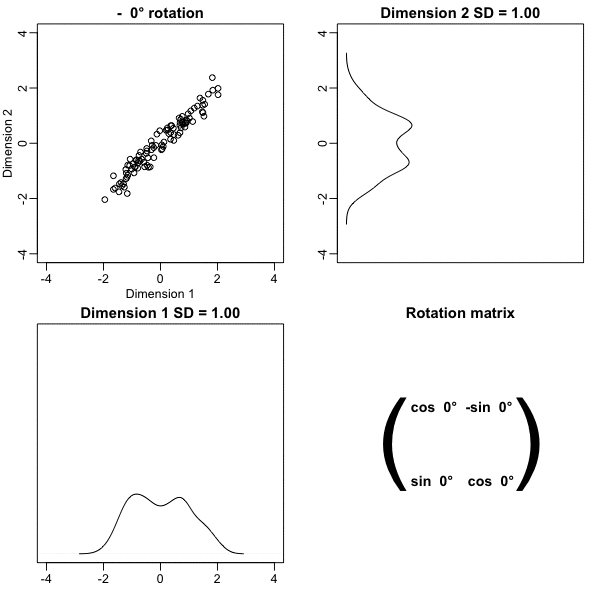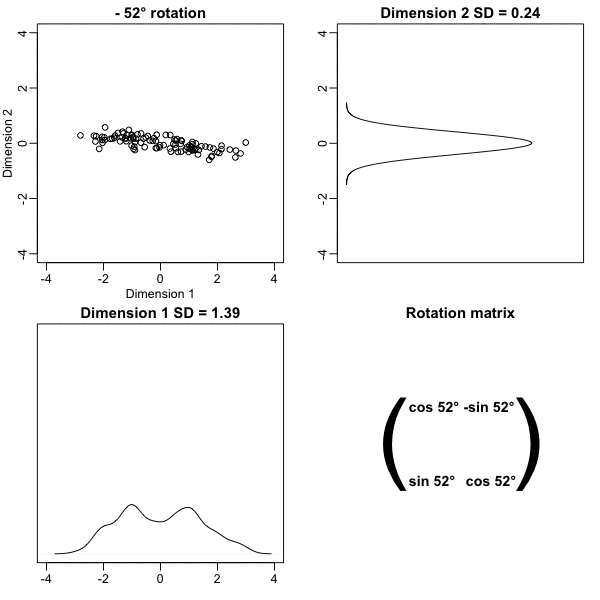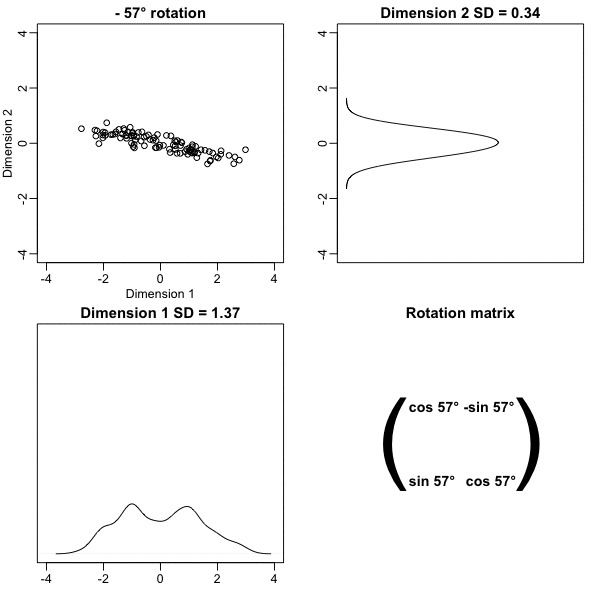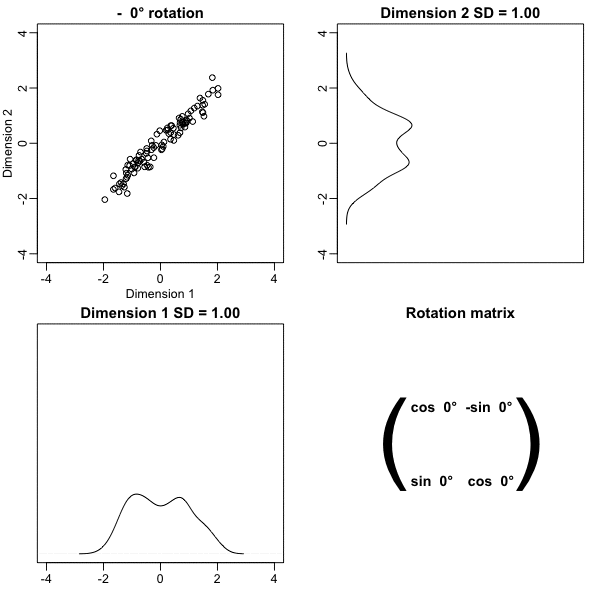
* In general, dimension reduction can be described as applying a rotation $A$ to a matrix with many columns $X$ that *moves* the information contained in $\mathbf{X}$ to the first few columns of $\mathbf{Z}=\mathbf{X}\mathbf{A}$.
* Then keeping just these few informative columns, reduces the dimension of the vectors contained in the rows.
* In our simplistic example we reduced the dimensions from 2 to 1, but the ideas extend to higher dimensions.
* In general, the first *principal component* (PC) of a matrix $\mathbf{X}$ is the linear orthogonal transformation that maximizes the variability of the first dimension.
* The function `prcomp` finds this transformation:
```{r}
pca <- prcomp(x)
pca$rotation
```
* Note that the first PC is almost the same as that provided by the $\cos(-45^{\circ}), \sin(-45^{\circ})$ we used earlier.
* The function `prcomp` returns the rotation needed to transform $\mathbf{X}$ so that the variability of the columns is decreasing from most variable to least (accessed with `$rotation`) as well as the resulting new matrix (accessed with `$x`).
* By default the columns of $\mathbf{X}$ are first centered (to change this change the `center` argument to `FALSE`).
* So, using the matrix multiplication shown above, we have that the following are the same (demonstrated by a difference between elements of essentially zero):
```{r}
#| eval: false
sweep(x, 2, colMeans(x))
pca$x %*% t(pca$rotation)
```
* The rotation is orthogonal which means that the inverse is its transpose.
* So we also have that these two are identical:
```{r eval=FALSE}
#| eval: false
sweep(x, 2, colMeans(x)) %*% pca$rotation
pca$x
```
* We can visualize these to see how the first component summarizes the data. In the plot below red represents high values and blue negative values, we learn why we call these weights and patterns:
```{r illustrate-pca-twin-heights, echo=FALSE, height = 5, out.width="70%"}
illustrate_pca <- function(x, flip=1,
pad = round((nrow(x)/2-ncol(x))*1/4),
cex = 5, center = TRUE){
rafalib::mypar(1,5)
## flip is because PCA chooses arbitrary sign for loadings and PC
colors = rev(RColorBrewer::brewer.pal(9, "RdBu"))
pca <- prcomp(x, center = center)
if(center) z <- t(x) - rowMeans(t(x))
cols <- 1:ncol(x)
rows <- 1:nrow(x)
image(cols, rows, z[,rev(1:ncol(z))], xaxt = "n", yaxt = "n",
xlab="", ylab="", main= "X", col = colors)
abline(h=rows + 0.5, v = cols + 0.5)
rafalib::nullplot(xaxt="n",yaxt="n",bty="n")
text(0.5, 0.5, "=", cex = cex)
z <- flip*t(pca$x)
image(cols, rows, z[,rev(1:ncol(z))], xaxt = "n", yaxt = "n",xlab="",ylab="", main= "Weights", col = colors)
abline(h=rows + 0.5, v = cols + 0.5)
rafalib::nullplot(xaxt="n",yaxt="n",bty="n")
text(0.5, 0.5, "x", cex = cex)
z <- flip*pca$rotation
nz <- cbind(matrix(NA, ncol(z), pad), z, matrix(NA, ncol(z), pad))
rows <- 1:ncol(nz)
image(cols, rows, nz[,rev(1:ncol(nz))], xaxt = "n", yaxt = "n", bty = "n", xlab="",ylab="", col = colors)
abline(h = pad+0:ncol(z)+1/2)
lines(c(ncol(z)/2+0.5,ncol(z)/2+1/2),c(pad,pad+ncol(z))+0.5)
text(ncol(z)/2+0.5, pad+ncol(z)+2 , expression(bold(Pattern^T)), font=2)
}
rafalib::mypar(1,1)
illustrate_pca(x, flip = -1)
```
* It turns out that we can find this linear transformation not just for two dimensions but for matrices of any dimension $p$.
* For a multidimensional matrix with $p$ columns, we can find a transformation that preserves distance between rows, but with the variance of the columns in decreasing order.
* The second column is the second principal component, the third column is the third principal component, and so on.
* As in our example, if after a certain number of columns, say $k$, the variances of the columns of $\mathbf{Z}_j$, $j>k$ are very small, it means these dimensions have little to contribute to the distance and we can approximate distance between any two points with just $k$ dimensions.
* If $k$ is much smaller than $p$, then we can achieve a very efficient summary of our data.
## Iris example
* The iris data is a widely used example in data analysis courses. It includes four botanical measurements related to three flower species:
```{r}
names(iris)
```
* If you print `iris$Species` you will see that the data is ordered by the species.
* Let's compute the distance between each observation.
* You can clearly see the three species with one species very different from the other two:
```{r}
x <- iris[,1:4] |> as.matrix()
d <- dist(x)
image(as.matrix(d), col = rev(RColorBrewer::brewer.pal(9, "RdBu")))
```
* Our predictors here have four dimensions, but three are very correlated:
```{r}
cor(x)
```
* If we apply PCA, we should be able to approximate this distance with just two dimensions, compressing the highly correlated dimensions.
* Using the `summary` function we can see the variability explained by each PC:
```{r}
pca <- prcomp(x)
summary(pca)
```
* The first two dimensions account for 97% of the variability.
* Thus we should be able to approximate the distance very well with two dimensions.
* We can visualize the results of PCA:
```{r illustrate-pca-twin-heights-iris, echo=FALSE, fig.height = 6, out.width="70%"}
rafalib::mypar()
illustrate_pca(x)
```
* And see that the first pattern is sepal length, petal length, and petal width (red) in one direction and sepal width (blue) in the other.
* The second pattern is the sepal length and petal width in one direction (blue) and petal length and petal width in the other (red).
* You can see from the weights that the first PC1 drives most of the variability and it clearly separates the first third of samples (setosa) from the second two thirds (versicolor and virginica).
* If you look at the second column of the weights, you notice that it somewhat separates versicolor (red) from virginica (blue).
* We can see this better by plotting the first two PCs with color representing the species:
```{r iris-pca}
data.frame(pca$x[,1:2], Species = iris$Species) |>
ggplot(aes(PC1, PC2, fill = Species)) +
geom_point(cex = 3, pch = 21) +
coord_fixed(ratio = 1)
```
* We see that the first two dimensions preserve the distance:
```{r dist-approx-4, message = FALSE, fig.height = 3, fig.width = 3, out.width="50%"}
d_approx <- dist(pca$x[, 1:2])
plot(d, d_approx); abline(0, 1, col = "red")
```
* This example is more realistic than the first artificial example we used, since we showed how we can visualize the data using two dimensions when the data was four-dimensional.
## MNIST example
* The written digits example has 784 features. Is there any room for data reduction? Can we create simple machine learning algorithms using fewer features?
* Let's load the data:
```{r}
library(dslabs)
if (!exists("mnist")) mnist <- read_mnist()
```
* Because the pixels are so small, we expect pixels close to each other on the grid to be correlated, meaning that dimension reduction should be possible.
* Let's try PCA and explore the variance of the PCs.
* This will take a few seconds as it is a rather large matrix.
```{r, cache=TRUE}
col_means <- colMeans(mnist$test$images)
pca <- prcomp(mnist$train$images)
```
```{r mnist-pca-variance-explained}
pc <- 1:ncol(mnist$test$images)
qplot(pc, pca$sdev)
```
* We can see that the first few PCs already explain a large percent of the variability:
```{r}
summary(pca)$importance[,1:5]
```
* And just by looking at the first two PCs we see information about the class.
* Here is a random sample of 2,000 digits:
```{r mnist-pca-1-2-scatter}
data.frame(PC1 = pca$x[,1], PC2 = pca$x[,2],
label = factor(mnist$train$label)) |>
sample_n(2000) |>
ggplot(aes(PC1, PC2, fill = label)) +
geom_point(cex = 3, pch = 21)
```
* We can also *see* the linear combinations on the grid to get an idea of what is getting weighted:
```{r mnist-pca-1-4, echo = FALSE, out.width="100%", fig.width=6, fig.height=1.75}
library(RColorBrewer)
tmp <- lapply( c(1:4,781:784), function(i){
expand.grid(Row = 1:28, Column = 1:28) |>
mutate(id = i, label = paste0("PC",i),
value = pca$rotation[,i])
})
tmp <- Reduce(rbind, tmp)
tmp |> filter(id < 5) |>
ggplot(aes(Row, Column, fill = value)) +
geom_raster() +
scale_y_reverse() +
scale_fill_gradientn(colors = brewer.pal(9, "RdBu")) +
facet_wrap(~label, nrow = 1)
```
* The lower variance PCs appear related to unimportant variability in the corners:
```{r mnist-pca-last,, echo = FALSE, out.width="100%", fig.width=6, fig.height=1.75}
tmp |> filter(id > 5) |>
ggplot(aes(Row, Column, fill = value)) +
geom_raster() +
scale_y_reverse() +
scale_fill_gradientn(colors = brewer.pal(9, "RdBu")) +
facet_wrap(~label, nrow = 1)
```
## Exercises
(@) We want to explore the `tissue_gene_expression` predictors by plotting them.
```{r, eval=FALSE}
dim(tissue_gene_expression$x)
```
We want to get an idea of which observations are close to each other, but the predictors are 500-dimensional so plotting is difficult. Plot the first two principal components with color representing tissue type.
(@) The predictors for each observation are measured on the same measurement device (a gene expression microarray) after an experimental procedure. A different device and procedure is used for each observation. This may introduce biases that affect all predictors for each observation in the same way. To explore the effect of this potential bias, for each observation, compute the average across all predictors and then plot this against the first PC with color representing tissue. Report the correlation.
```{r}
avg <- rowMeans(tissue_gene_expression$x)
pca <- prcomp(tissue_gene_expression$x)
data.frame(PC1 = pca$x[,1], avg = avg, tissue = tissue_gene_expression$y) |>
ggplot(aes(PC1, avg, color = tissue)) +
geom_point() +
ggtitle(paste("The correlation is", cor(pca$x[,1],avg)))
```
(@) We see an association with the first PC and the observation averages. Redo the PCA but only after removing the center.
(@) For the first 10 PCs, make a boxplot showing the values for each tissue.
(@) Plot the percent variance explained by PC number. Hint: use the `summary` function.